Example 1: Fill Indian Ocean with land
!!! This page is under construction !!!
In this example, you can learn how to modify the default continental geometries in an uncoupled case (either an F-case with atmosphere and land, or an I-case with land only) and specify the land surface properties in the modified continental regions. First see the information on mesh and fsurdat file as you will be modifying these files in this example. If you are using CESM and the CESM tag is located in the directory \$CESM, then the tools that you will be using are located in \$CESM/components/clm/tools/modify_input_files. The following describes how to use these tools within \$CESM, but if you are using CTSM instead, then these tools will be located at \$CTSM/tools/modify_input_files, where \$CTSM is the location of your CTSM tag. These tools can be run from the command line, or alternatively, they can be run interactively from the VisualCaseGen GUI.
To fill in the Indian Ocean with land, there are two primary things that need to be altered:
- The fsurdat file which specifies the land surface properties. See more information on the fsurdat file and how to find the default for your grid resolution and compset here.
- The mesh file which controls the land mask. See more information on the mesh file and how to find the default for your grid resolution and compset here.
Step 1: Setting up your case
Set up the case using the default continental geometry. In this example, we'll use the "FHIST" compset (i.e., prescribed time varying historical Sea Surface Temperatures and forcings with satellite phenology used in the land) and we'll use the grid "f09_f09_mg17" which is a nominal 1 degree grid.
cd $CESM/cime/scripts
./create_newcase --case path_to_case/case_fillIO.001 --compset FHIST --res f09_f09_mg17where "path_to_case" refers to the refers to the file path where we'd like to store the case directory and case_fillIO.001 is our chosen case name for "fill Indian Ocean".
Step 2: Modify the fsurdat file
To do this we make use of the fsurdat modifier tool located at \$CESM/components/clm/tools/modify_input_files/fsurdat_modifier which reads in information from the file \$CESM/components/clm/tools/modify_input_files/modify_fsurdat_template.cfg.
Step 2.1: Provide the inputs to the fsurdat modifier tool.
It is recommended that you make a copy of modify_fsurdat_template.cfg for each new experiment. We've called the experiment fillIO so we'll call out new .cfg file "modify_fsurdat_fillIO.cfg i.e.,
cd $CESM/components/clm/tools/modify_input_files/
cp modify_fsurdate_template.cfg modify_fsurdat_fillIO.cfgIn modify_fsurdat_fillIO.cfg we'll set all the parameters to fill in the Indian ocean with land and set our chosen land surface type.
The region that you'd like to change from ocean to land can be specified as a mask that can be read in from a file by the fsurdat modifier tool. It requires an input netcdf file containing the following variables:
| landmask(lsmlat,lsmlon) | landmask containing 1's for land, 0's elsewhere |
| mod_lnd_props(lsmlat,lsmlon) | mask where the surface properties will be altered (1's for modification, 0's elsewhere) |
| lats(lsmlat) | the latitudes of the grid |
| lons(lsmlon) | the longitudes of the grid |
In this example where the default continental geometry is being modified, you can first find the default landmask and modify that (see the instructions for "Finding the default land mask" here) and then modify this to add or remove land, as required. In this case, we are adding land grid points to the Indian Ocean. This example script uses the default landmask for an FHIST case in CESM2 at f09_f09_mg17 resolution and produces the masks required for the fsurdat modifier tool as depicted in Figure 1.
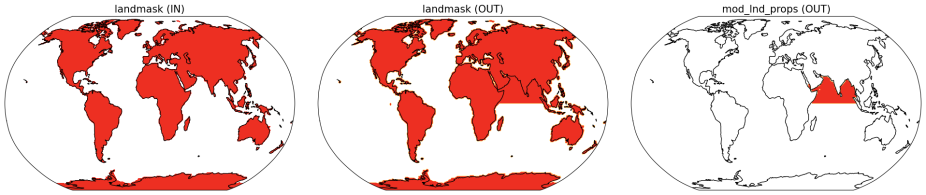
Fig 1: (left) Input land mask, (middle) output land mask with the Indian Ocean filled, (right) output mod_lnd_props indicating regions where the land surface properties will be modified. Red colors indicate values equal to 1.
We have called the file containing these new masks "mask_fillIO.nc" and we will now put it in a subdirectory \$CESM/components/clm/tools/modify_input_files/myinputs/masks. Now we need to provide the following inputs to our modify_fsurdat_fillIO.cfg file.
| fsurdat_in | the default fsurdat file for the grid/compset you want to use (see here for how to find the default fsurdat file) |
| fsurdat_out | a path where you'd like to put your modified fsurdat file |
| idealized | if True, idealized land surface properties can be specified where mod_lnd_props=1 |
| landmask_file | a file containing the landmask and mod_lnd_props variables that you have just made |
| dom_pft | a number that specifies the plant functional type you'd like to use where mod_lnd_props=1 |
| lai | Twelve values for the leaf area index for each month of the year |
| sai | Twelve values for the stem area index for each month of the year |
| hgt_top | Twelve values for the height of the bottom of the canopy for each month of the year |
| hgt_bot | Twelve values for the height of the bottom of the canopy for each month of the year |
| include_nonveg | if True, non-vegetated land units remain unchanged e.g., glacier, wetland, lake, urban |
In this example, we set the following
fsurdat_in = $FSURDAT_ORIG
fsurdat_out = ./myinputs/fsurdat/fsurdat_fillIO.nc
idealized = True
landmask_file = ./myinputs/masks/mask_fillIO.nc
dom_pft = 14
lai = 3 3 3 3 3 3 3 3 3 3 3 3
sai = 1 1 1 1 1 1 1 1 1 1 1 1
hgt_top = 1 1 1 1 1 1 1 1 1 1 1 1
hgt_bot = 0.5 0.5 0.5 0.5 0.5 0.5 0.5 0.5 0.5 0.5 0.5 0.5
include_nonveg = Truewhere $FSURDAT_ORIG refers to the default fsurdat file for the compset and resolution, that we can find following the instructions here. In this example, we have chosen a plant functional type (PFT) of 14, which corresponds to c4-grass. The PFT numbers and names may change depending on which model version or configuration you're using, but you can look in the "paramfile" for the default case to find out the numbers that correspond to different PFTs for your case (see section 2.2 here). We have also set the leaf area index to 3 for each month of the year and the stem area index to 1 for each month of the year, the top of the canopy to 1 for each month of the year and the bottom height of the canopy to 0.5 for each month of the year. These affect radiation and momentum transfer within the plant canopy, but note that they only need to be specified when using satellite phenology mode, otherwise the model would be simulating these.
Step 2.2: Running the fsurdat modifier tool
In order to run the fsurdat modifier tool, you'll need to activate the conda environment "ctsm_pylib". If you don't yet have the ctsm_pylib environment, you'll need to create it and activate it by running the following (note that if you're on the NCAR machines you may need to also load the conda module)
./$CESM/components/clm/py_env_create
conda activate ctsm_pylibNow you can run the fsurdat_modifier tool on your input file modify_fsurdat_fillIO.cfg i.e.,
cd $CESM/components/clm/tools/modify_input_files
./fsurdat_modifier modify_fsurdat_fillIO.cfgThis can take up to several hours and you should find that a modified fsurdat file has been created at the location you gave to fsurdat_out. This is your new fsurdat file that you'll use in your case (see step 4).
Step 3: Modifying the mesh file
The mesh_mask_modifier tool can be used to modify the mesh file to represent the new landmask for your case. This is done in a similar manner to what is done in step 2 above for the fsurdat file but instead uses the modify_mesh_template.cfg and the mesh_mask_modifier files. Make a copy of the template file for the fill Indian Ocean case:
cd $CESM/tools/modify_input_files
cp modify_mesh_template.cfg modify_mesh_fillIO.cfgIn modify_mesh_fillIO.cfg, the following variables will need to be set:
| mesh_mask_in | points toward the default mesh file for your case |
| mesh_mask_out | points toward the location you'd like to store your modified mesh file |
| landmask_file | the location of the file that contains your modified landmask |
| lat_dimname | the name of the latitude dimension in landmask_file (lsmlat in this example) |
| lon_dimname | the name of the longitude dimension in landmask_file (lsmlon in this example) |
| lat_var | the name of the latitude variable in landmask_file (lats in this example) |
| lon_var | the name of the longitude variable in landmask_file (lons in this example) |
In this example, we set the following
mesh_mask_in = /glade/p/cesmdata/cseg/inputdata/share/meshes/fv0.9x1.25_141008_polemod_ESMFmesh.nc
mesh_mask_out = ./myinputs/mesh/mesh_fillIO.nc
landmask_file = ./myinputs/masks/mask_fillIO.nc
lat_dimname = lsmlat
lon_dimname = lsmlon
lat_varname = lats
lon_varname = lonswhere "mesh_mask_in" was determined using the steps in section 1.2 here and we have set the landmask_file to the file we generated in step 2.1 above. Make sure you still have the ctsm_pylib environment activated and now the mesh modifier tool can be run by
cd $CESM/components/clm/tools/modify_input_files/
./mesh_mask_modifier modify_mesh_fillIO.cfgThis can take a few minutes.
Step 4: Configure the case
We have set up the case in Step 1, but now we need to point this case to our modified fsurdat and mesh files. Our case is located in path_to_case/case_fillIO.001. So we can go in there an set up the case
cd path_to_case/case_fillIO.001
./case.setupWe can point this case to our new fsurdat file by adding the following line to user_nl_clm within our case directory
fsurdat='$CESM/components/clm/tools/modify_input_files/myinputs/fsurdat/fsurdat_fillIO.nc'If you are running a case with time varying land use, such as an FHIST case, the following will also need to be added to user_nl_clm
check_dynpft_consistency=.false.Upon initialization, a check is performed for consistency between the surface dataset and the land use change input and in this case they will not be consistent and the above namelist setting makes the model skip this check.
We can point this case to our new mesh file by modifying the variable MASK_MESH in env_run.xml.
./xmlchange MASK_MESH=$CESM/components/clm/tools/modify_input_files/myinputs/mesh/mesh_fillIO_002.nc We will also need to enforce that CLM starts from a cold start since the default initial conditions will not contain any land surface properties over the Indian ocean region. This can be done using the following xmlchange command
./xmlchange CLM_FORCE_COLDSTART=onStep 5: build and submit the case
Now you can build your case
./case.buildNote that if you're on the NCAR computer, you'd need to instead use
qcmd -- ./case.buildThen submit your case using
./case.submit